CYPs are mighty synthesis machines
This is an insightful article by a group at University of Dundee and Hypha that illustrates, as the title says, late-stage functionalization using P450 enzymes. It is surprising to me that this idea and implementation of the idea has not caught on more widely. As a bit of background, in a paper that recently published, I outlined that more than 40% of FDA oral approved drugs (2020-2024) are metabolized by CYP3A4, and a large portion metabolized by other CYP enzymes (which is no surprise if you work in the field) (Figure 1). I characterized a list of ~14 drugs that had active metabolites, among which several were hydroxylations on alkyl side chains, which in some cases may be hard to get to otherwise (in the case of gepirone).

Now, with that in mind, let’s consider the paper from Dundee and Hypha. The team illustrated the use of commercially available enzyme screening panels, such as Codexis MicroCYPs and the kit used herein, Hypha Discovery’s PolyCYPs. More importantly, this functionalization incorporates polar and hydrophilic substituents, which is often an Achilles heel of many drug discovery programs, where off-target and poor drug-like properties are correlated to high lipophilcity.
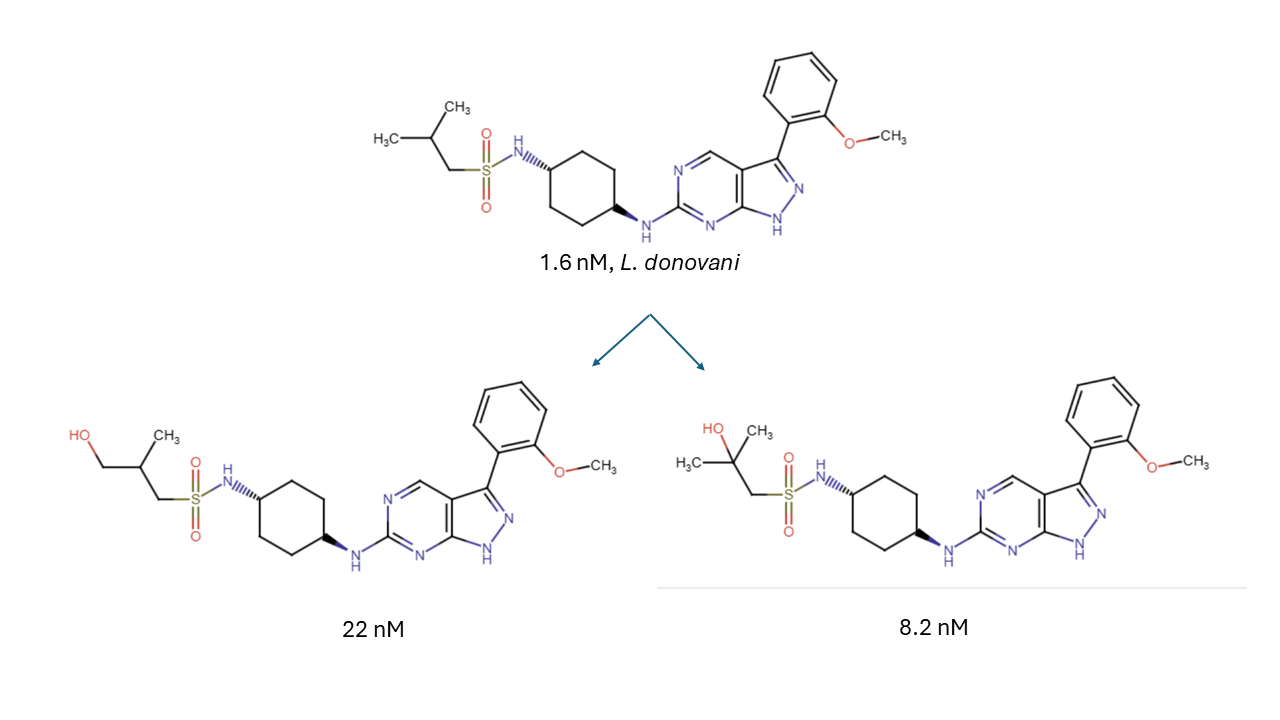
The experiment began by screening twelve structurally diverse lead compounds in the CYP panel from in-house antiparasitic drug discovery programs at concentrations of 0.1 mg/mL, a concentration which reduces risk of enzyme inhibition effects. The PolyCYPs panel consisted of twenty-three PolyCYPs enzymes, five FMO enzymes, and one AO enzyme. Cases where successful products were identified, the reactions were scaled up (volumes to between 30 and 250 mL, at a 0.1 mg/mL substrate using either lyophilized scale-up vials or fresh enzyme extract from E. coli-derived processing). The authors obtained twenty-three products between 0.66 mg–11.16 mg yield following purification. They then screened these new products against a panel of pathogens (e.g. Schistosoma mansoni, Leishmania donovani, Trypanosoma cruzi, and Trypanosoma brucei). Several novel hits were obtained, illustrated below.

To me, this approach should be more widely examined as a way to incorporate novel chemical diversity with hydrophilic substitutions.

Comments are closed